Producing Graphics From MEIS Crystals
This page describes a method for producing semi-decent graphical representations of MEIS crystals. without too much “drawing”. I refined this method in order to rapidly produce ball and stick diagrams whilst making the minor corrections to my thesis. This page will probably be of greatest interest to the University of York Surface Physics Group, where that doctoral work was performed. If you have any queries you can email me.
The Method
To produce a fairly decent ball and stick diagram I went through the following steps. The software required is described at the end.
- Produce your crystal. If your working on MEIS you probably already have one. If not then use XVEGAS to make one, or do it by hand, they’re simple text files.
- Grow your crystal. MEIS crystals tend to contain only one unit cell. You’ll probably want more than that in your graphic. Open XVEGAS and select the Basis menu, then the Crystal Menu. Open a crystal file. Select grow. Check the x or y radio button and enter the salient information (note repeat distances for x and y are displayed at the top of the main crystal window—don’t ask me why it doesn’t grab this information for you). Repeat as desired (you might have to iterate through the steps to get a crystal as big as you want). Group the crystal and then Save it (Note: The software will add .crs to the end even if you specify an extension for the filename).
- Now, in a moment we’re going to convert the MEIS crystal file into another format. However, the software to do this seems very picky about the exact formatting of the input text file and the file outputted by XVEGAS doesn’t seem to be quite the right format (sigh). You’ll be wanting to run my x2magic script on the crystal file (see software) to produce a new crystal file.
- Take the new crystal file just produced and run it through Vegas Magic. Convert it to a PDB file
- Finally the PDB file just produced can be loaded into JMol. There you can rotate the crystal around, zoom and quite usefully delete atoms (this also deletes the associated bonds, surprisingly enough). You can capture the view in a number of image formats for manipulation in the graphics package of your choice (job done!). The view can also be exported as a .pov file for rendering in POVRay.
Additional Notes
The display of the atoms in JMol is controlled by the file jmol_atomtypes.txt , located (under XP) in Documents and Settings\user\.jmol It’s basically lines of tab separated columns. Note that not every atom is included in the default file, (notably the rare earths aren’t). The columns are, left to right, the atom type, base atom (not sure what this does), atomic number, atomic mass, van der Waals radius, covalent radius, and the RGB representation fo the rendering colour (in base 10, i. e. 0 to 255). Note that the wan der Waals radius controls the atom size. You can make atoms up with false symbols but notice that despite appearances, symbols are not case sensitive, the program just uses the first one in the file. You could therefore edit the PDB crystal file to change certain atoms to this imaginary type in order to render them distinctly (I think changing the type in the MEIS crystal file causes problems at some stage so I advice against it).
Software
The following software was used in the stages above.
- XVEGAS
- XVEGAS is the TCL/TK simulation software (based on the original vegas codes) produced at Warwick. Your crystal may well originate from this. In any case it is useful in order to grow the crystal (the MEIS crystal in general contains only one unit cell, but in a graphic we probably want more than one). Unfortunately Warwick’s web pages aren’t very forthcoming on the subject. Check my resources page.
- JMol
- Jmol is a free, open source molecule viewer for students, educators, and researchers in chemistry and biochemistry. It is cross-platform, running on Windows, Mac OS X, and Linux/Unix systems. It’s a Java application (and also a Java applet which may be embedded in web pages). It should be possible to substitute RasMol here, but I didn’t.
- Vegas Magic
- Converts MEIS crystals into PDB format (or into Alchemy format). Thanks to Nick Kaijaks.
- x2magic
- A c++ program I hacked in desperation to try and produce figures for thesis corrections in a reasonable time. See my resources page for more details.
Finally, as an example, part of a figure from my thesis, showing iron silcide. Notice how the additional silicon adatoms have been specially rendered in a bright yellow.
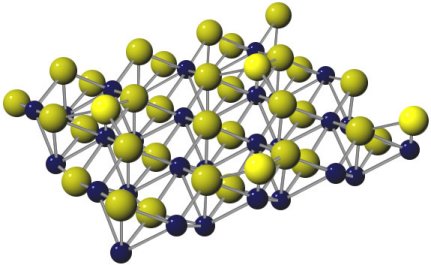
Example Graphic
A graphic produced by the procedure
Comments and Pings